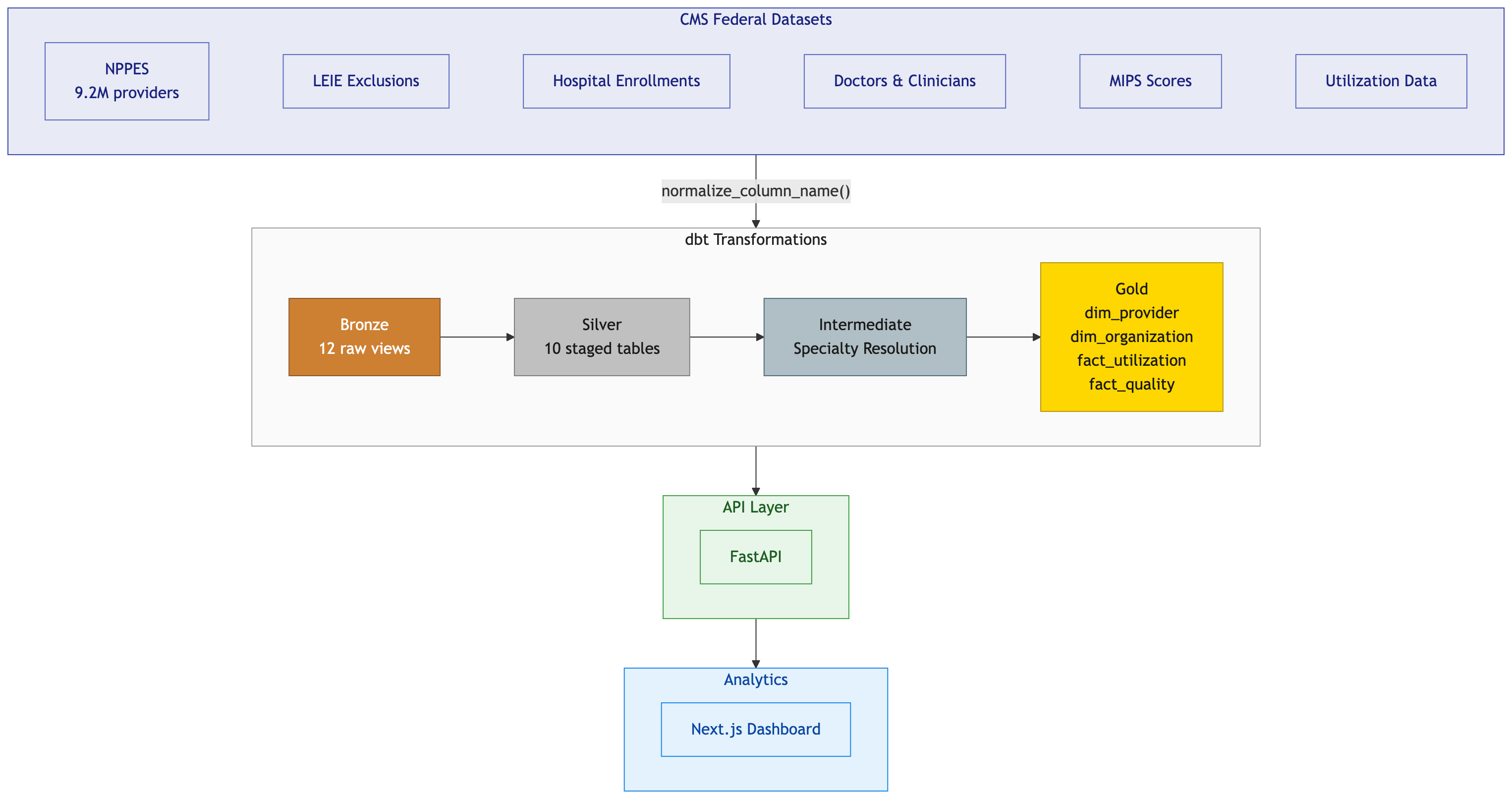
What Broke When I Pointed My dbt Project at Real Data
I built a healthcare provider data platform in a weekend using Claude Code. Mock data generators, a full dbt medallion architecture, 449 features with automated tests. Everything passed. DuckDB humming along locally, every model green.
Then I pointed it at real CMS data on BigQuery and watched the whole thing fall apart.
Not in one dramatic failure. In dozens of small ones. SQL that worked fine on DuckDB threw errors on BigQuery. Column names that were perfectly valid in one engine were illegal in the other. Data that my mock generators said would never exist showed up in the hundreds of thousands. I spent more time fixing what I thought was done than I spent building it in the first place.
Here’s what I learned.
Mock Data Is a Confidence Trap
My mock data generators were thorough. I modeled NPPES with all 330 columns. I got the Luhn checksum right on NPI numbers. I matched real CMS distributions for exclusion types, hospital ownership, and charge-to-payment ratios. I even handled the different date formats across datasets (NPPES uses MM/DD/YYYY, LEIE uses YYYYMMDD, because of course they do).
All 449 tests passed. I felt great about it.
Then real NPPES data arrived. 9.2 million rows instead of 25. And immediately, five distinct data quality issues surfaced that my mock data never anticipated.
17,738 records had date strings in the taxonomy code field. Not taxonomy codes. Dates. Like 03/27/2006 sitting in a column that should contain something like 207R00000X. A data entry error at the CMS source that has apparently persisted for years. My mock generator would never produce this because I explicitly generated valid taxonomy codes. The real world doesn’t care about your schema.
1.9 million organizations had no taxonomy code at all. My mock data always populated taxonomy for every provider. Turns out taxonomy is optional for Entity Type 2 (organizations) per NPPES rules. Nearly two million test failures because I assumed a field would be populated based on what made sense to me rather than what the spec actually says.
51,975 MIPS scores had duplicate NPIs. I assumed NPI was the primary key for MIPS data. It’s not. The source file is keyed on NPI plus Org_PAC_ID, because a provider affiliated with multiple groups gets a separate score for each. My uniqueness test exploded.
Two Order/Referring NPIs appeared twice with slightly different name spellings. ANABELL versus ANABEL. CMS applied a name correction without removing the prior row. Two records. That’s it. But two is enough to break a uniqueness constraint.
334,000 rows were ghost records. Valid NPI, every other field empty. These are deactivated providers that CMS keeps in the file indefinitely. 3.6% of the entire dataset is just… hollow shells. My entity type mapping produced “Unknown” for these because the entity type was an empty string, not NULL. A distinction that matters exactly once and costs you hours when you discover it.
Every single one of these passed on mock data. Not because my tests were bad. Because my mock data was too clean. It modeled what CMS data should look like, not what it actually looks like.
The fix for each issue wasn’t complicated. Regex validation for taxonomy codes. Scoping the specialty test to individuals only. ROW_NUMBER dedup for MIPS. GROUP BY with most-permissive eligibility for the name variants. A WHERE clause filtering empty entity types. Simple stuff. The hard part was finding them.
I ended up writing formal Data Quality Decision Records (DQDRs) for each issue, documenting the investigation, the root cause, and the fix. That process turned out to be more valuable than the fixes themselves because it forced me to think about what other assumptions I was making that real data would eventually violate.
DuckDB and BigQuery Speak Different Languages
dbt is supposed to abstract away database differences. Write your SQL once, run it anywhere. That’s the promise.
The reality is more nuanced. dbt handles the basics well. ref() works the same everywhere. Materializations translate cleanly. But the moment you write any actual SQL beyond SELECT and WHERE, you’re in dialect territory.
I developed everything on DuckDB locally. Fast, zero setup, embedded in the Python process. Perfect for development. Then I deployed to BigQuery and discovered how many DuckDB-isms had crept into my SQL.
Some differences were obvious. DuckDB uses ::VARCHAR for casting. BigQuery wants CAST(... AS STRING). Easy enough, switch to dbt.safe_cast() and move on.
Some were subtle. TRY_STRPTIME(field, format) in DuckDB becomes SAFE.PARSE_DATE(format, field) in BigQuery. Same function, same purpose, but the argument order is flipped. That’s the kind of thing that compiles fine and silently produces wrong results if you get it backwards.
Some were structural. DuckDB supports DISTINCT ON (column) for deduplication. BigQuery doesn’t have it. You need the ROW_NUMBER() OVER(PARTITION BY ...) pattern instead. DuckDB lets you define inline tables with VALUES (...) AS t(col1, col2). BigQuery requires you to spell it out as chained UNION ALL SELECT statements. I ended up writing Jinja loops to generate these.
The list kept growing. CONCAT_WS versus ARRAY_TO_STRING. SPLIT_PART versus SPLIT()[OFFSET()]. REGEXP_MATCHES versus REGEXP_CONTAINS. The modulo operator % versus MOD(). Each one is a five-minute fix in isolation. Multiply by dozens of occurrences across a full dbt project and you’ve lost a day.
Then there were the truly annoying ones. BigQuery requires typed NULLs in UNION ALL queries. You can’t just write NULL. You need CAST(NULL AS STRING). DuckDB doesn’t care. I had a dim_organization model that worked perfectly locally and threw a type inference error on BigQuery because of a bare NULL in one branch of a UNION.
Reserved words bit me twice. I used grouping as a column name in my taxonomy staging model. Fine in DuckDB. Reserved in BigQuery. I used source as a CTE name in my MIPS model. Same story. Both required renaming, which meant updating every downstream reference.
I ended up building a cross_db.sql macro file with adapter-aware wrappers for every function that differed. It works, but it’s the kind of infrastructure you wish you didn’t need.
The lesson here isn’t “don’t use DuckDB for development.” DuckDB is fantastic for development. The lesson is that if you’re targeting BigQuery (or Snowflake, or Redshift) in production, run your tests against that engine early. Not after a weekend of local-only development. Not after you’ve declared all features complete. Early.
There’s a practical gap here too. With a dataset like NPPES at 9.2 million rows, you can’t really analyze data quality from a CSV on your laptop. You need it in a warehouse first. But the naive approach of “load it raw, then build transformations” skips a critical step. Before you write a single staging model, spend time with the raw data. Run distribution queries. Look for NULLs where you don’t expect them, values that don’t match the data dictionary, duplicate keys. Use an LLM to help scan for outliers if the dataset is too wide to eyeball (330 columns is too wide to eyeball). The raw ingestion layer is where you learn what the data actually looks like, and that knowledge should inform your transformation logic, not the other way around.
The other thing I’d add to the process: BigQuery validation in CI. If I’d had a GitHub Action running dbt build against a BigQuery dataset on every push, each dialect issue would have surfaced as a single failing commit instead of a wall of errors after the fact. The cost of a small BigQuery CI dataset is trivial compared to the cost of debugging dozens of cross-engine issues in one sitting.
The Column Name Normalization Saga
This one deserves its own section because it cost me more time than any SQL dialect issue and I went through two failed approaches before landing on the right one.
CMS publishes CSV files with column headers like Provider Last Name (Legal Name), SUBGROUP - GENERAL, and Healthcare Provider Primary Taxonomy Switch_1. Spaces, parentheses, hyphens, slashes, mixed case. The full NPPES file has 330 of these columns.
DuckDB handles this fine. It preserves the original names and lets you reference them with double quotes. "Provider Last Name (Legal Name)" works exactly as you’d expect.
BigQuery’s CSV autodetect takes a different approach. It sanitizes column names by replacing special characters with underscores. But it does this inconsistently. Spaces become underscores. Parentheses become underscores. Sometimes you get double underscores, sometimes you don’t. Columns that start with numbers get a leading underscore. And here’s the really fun one: columns containing only Y and N values get inferred as BOOL instead of STRING.
My first approach was to just handle it in the SQL. Double-quote everything for DuckDB, use the sanitized names for BigQuery. This worked for about three models before the inconsistencies became unmanageable.
My second approach was a Jinja macro called col(). The idea was clean. Write `` in your model and the macro would produce the right column reference for whichever adapter you were running on. For DuckDB, it would output the double-quoted original name. For BigQuery, it would apply the same sanitization rules BigQuery uses.
I updated all ten silver models to use it. It worked for the first few CSVs. Then I hit a column where BigQuery’s sanitization produced a different result than my macro predicted. Fixed that. Hit another one. Fixed that. A column starting with a digit got a leading underscore that my macro didn’t account for. Fixed that. A Y/N column became BOOL. Fixed that.
It was whack-a-mole. Every CSV file had some new edge case in how BigQuery chose to mangle its column names. The macro grew more complex with each fix, and I had zero confidence it would handle the next file correctly.
The col() macro lasted about two hours before I ripped it out.
The third approach was the right one, and in retrospect it was obvious. Don’t try to adapt to whatever name each engine gives your columns. Normalize the names yourself at the point of ingestion, before the data ever reaches dbt.
I wrote a single Python function called normalize_column_name(). It lowercases everything, replaces spaces and special characters with underscores, collapses multiple underscores, and strips leading/trailing underscores. This function runs in two places: the DuckDB loader script that reads CSVs locally, and the BigQuery schema generator that produces JSON table definitions for external tables.
Both engines now see identical column names. provider_last_name_legal_name. subgroup_general. healthcare_provider_primary_taxonomy_switch_1. Plain lowercase snake_case everywhere. The silver models dropped all the quoting and macro calls and just reference columns by their normalized names.
The commit that implemented this touched 33 files and changed over 3,000 lines. That includes JSON schema definitions for every CMS source table (the NPPES schema alone is 1,652 lines because of those 330 columns). It was a big change. It was also the last time I had a column name problem.
The takeaway is something I should have recognized from the start. When you have a boundary between systems that don’t agree on conventions, normalize at the boundary. Don’t push the translation logic downstream into every consumer. Don’t build clever abstractions that try to bridge the gap at query time. Just make the data look the same before it enters your system.
What I’d Do Differently
If I were starting this project over, three things would change.
First, I’d test against the real target engine from day one. DuckDB for local development is great, but I’d set up a BigQuery dataset in the first week and run dbt test against it after every batch of model changes. Every dialect issue I found would have surfaced immediately instead of accumulating into a wall of failures.
Second, I’d seed my mock data with garbage. Not just valid data that matches the schema. Intentionally dirty data that matches what real-world sources actually produce. Empty strings where you expect NULLs. Date values in non-date columns. Duplicate keys with slight spelling variations. Deactivated records with hollow fields. If I’d included even a handful of these edge cases in my test data, four of my five data quality issues would have been caught before I ever touched real data.
Third, I’d normalize column names before writing a single dbt model. The column naming problem was entirely predictable. CMS files have messy headers. Different databases handle messy headers differently. The solution (normalize at ingestion) is a one-time investment that eliminates an entire category of problems. I spent time on two wrong approaches first because I kept trying to solve it closer to where the pain was felt instead of where the pain originated.
The Broader Pattern
Every one of these problems has the same shape. Something works in a controlled environment. You build confidence. You ship it. Then the uncontrolled environment reveals assumptions you didn’t know you were making.
Mock data assumes fields are populated. Real data has gaps. Local databases assume standard SQL. Production databases have dialects. Your column naming strategy assumes consistent behavior. Different engines have different opinions.
The fix is always the same too. Push your validation closer to reality. Test against real engines, real data shapes, and real edge cases. And when you find a boundary between two systems that disagree, normalize at the boundary instead of translating downstream.
None of this is revolutionary. But I had to learn it the expensive way, one broken pipeline at a time.
If you’re building healthcare data pipelines and want to compare war stories, connect with me on LinkedIn.